| E-mail ID : info@iamg.in |
| E-mail ID : info@iamg.in |
Online Submission |
| Click Here For Online Submission |
| Instructions for authors |
Genetic Clinics |
| Editorial board |
Get Our Newsletter |
| Subscribe |
Send Your Feedback |
| Feedback Form |
About Us |
| IAMG |
GeNeViSTA
| S. No. | Gene | Processed /Non processed pseudogene | Disease |
| 1 | IDS | Non processed | Hunter syndrome |
| 2 | GBA | Non processed | Gaucher disease |
| 3 | NCF1 | Non processed | Chronic granulomatous disease |
| 4 | PKD1 | Non processed | Autosomal dominant polycystic kidney disease |
| 5 | IGLL1 | Non processed | B-cell deficiency |
| 6 | ABCC6 | Non processed | Pseudoxanthoma elasticum |
| 7 | CYP21A2 | Non processed | Congenital adrenal hyperplasia |
| 8 | FOLR1 | Non processed | Neural tube defects |
| 9 | SBDS | Non processed | Shwachman-Diamond syndrome |
| 10 | VWF | Non processed | Type 3 von Willebrand disease |
| 11 | CRYBB2 | Non processed | Congenital adrenal hyperplasia |
Hunter syndrome (MPS II) is caused by mutations in the IDS gene [Timms et al., 1995]. Pseudogene of IDS known as IDS2 or IDSP1 shows 80kb similarity with the functional IDS gene. Pseudogene IDS2 is 96% homologous in exon 2, and intron 2, 3 and 7 of the transcribed IDS gene and 100% similar to exon 3 of IDS gene. The presence of pseudogene can result in recombination with the IDS gene. IDS-IDS2 undergoes homologous recombination [Lualdi et al., 2005] and thus is involved in the formation of large complex genomic/genetic rearrangements, deletions etc. which comprise about ˜13% of the patients with Hunter syndrome. Similar type of gene conversion events are known to occur in patients with Gaucher disease [Tayebi et al., 2003]. Recombinant alleles of GBA gene for Gaucher disease arise due to presence of the pseudogene GBAP 16 kb downstream of the functional gene. This pseudogene has 96% exonic sequence homology with GBA, with the region between intron 8 and 3’ UTR being >98% homologous. This promotes gene conversion and gene fusion events by non-reciprocal and reciprocal recombination. These events are most commonly seen in the exon 9–11 region. At least 10 GBA recombinants have been reported which comprise as many as 20–30% of alleles in some populations [Koprivica et al., 2000].
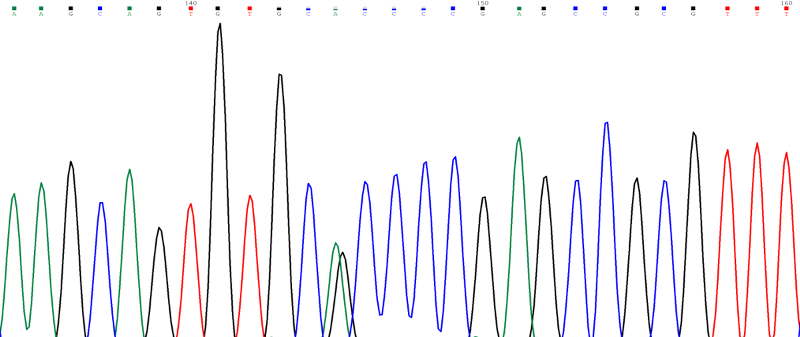
Case Study: Sanger sequencing of the IDS gene in a male patient with Hunter syndrome (detected by deficient enzyme assay) revealed a heterozygous variant p.Ala85Thr (alanine to threonine at amino acid 85) caused by a G to A substitution at nucleotide position c.253 in exon 3 of IDS (Figure 4). Since a male has only one X chromosome and the IDS gene is present on the X chromosome, we expect the patient to be hemizygous for the variant with a single peak on Sanger sequencing. The heterozygous peak in this chromatogram was seen because the mutant allele came from gene and the normal allele came from the pseudogene. Thus, it is important to design primers for polymerase chain reaction (PCR) in such a way that only the gene is amplified and the pseudogene is not amplified for sequencing of genes where pseudogenes are known to be present in the genome. Similar problem is faced in interpretation of results in next generation sequencing assays like exome sequencing and hence this should be kept in mind while interpreting the results.
In molecular diagnostics for genetic diseases, it is important that the sequencing information should come from the gene and not from the pseudogene. Hence, modifications need to be done to identify a gene. This can be done in two ways:
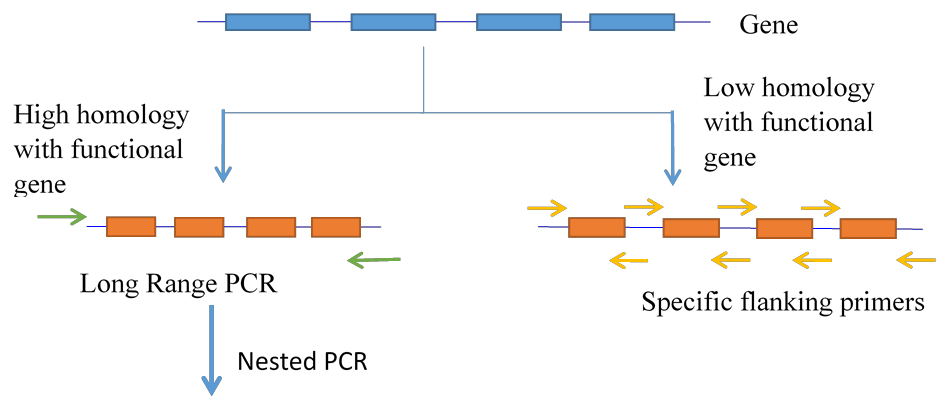
1. If a pseudogene is highly homologous to the functional gene and there are no differences in intronic regions, then the strategy employed is Long Range PCR followed by nested PCR. In this method a long-range PCR assay is designed using primers designed in such a way that the 3’ end of the primer falls on a single nucleotide change between gene and pseudogene found nearest to the gene. This is followed by a long PCR using special DNA Polymerase and then the PCR product is used as a template for subsequent PCR and sequencing using flanking primers for each exon. The long PCR ensures that only the gene is sequenced for interpretation of results (Figure 5).
2. If there are multiple differences in intronic regions of the gene and the pseudogene then flanking primers for each exon can be designed in such a way that the 3’ end of primer falls on a single nucleotide change. Thus, there is no need of long PCR in such cases (Figure 5).
Pseudogenes are essential parts of gene regulation. Understanding the mechanism of pseudogene actions is likely to help researchers to solve several essential biochemical pathways. Pseudogenes might be functional and participate in gene expression and molecular mechanisms of gene interactions. Pseudogenes can lead to genetic diseases due to gene conversion and they also pose a problem in genetic diagnostics.
1. Koprivica V, et al. Analysis and classification of 304 mutant alleles in patients with type 1 and type 3 Gaucher disease Am J Hum Genet 2000; 66:1777-1786.
2. Lualdi S, et al. Characterization of iduronate-2-sulfatase gene–pseudogene recombinations in eight patients with Mucopolysaccharidosis type II revealed by a rapid PCR-based method. Hum Mutat 2005; 25: 491-497.
3. Tayebi N, et al. Reciprocal and nonreciprocal recombination at the gluco cerebrosidase gene region: implications for complexity in Gaucher disease. Am J Hum Genet 2003; 72: 519-534.
4. Timms KM, et al. 130 kb of DNA sequence reveals two new genes and a regional duplication distal to the human iduronate-2-sulfate sulfatase locus. Genome Res 1995; 5: 71-78.
5. Tutar L, et al. Involvement of miRNAs and pseudogenes in cancer. Methods Mol Biol 2018; 1699: 45-66.
| Abstract | Download PDF |